Steve Gygi and Wade Harper (Harvard, Cell Biology) and Susan Celniker (Berkeley)
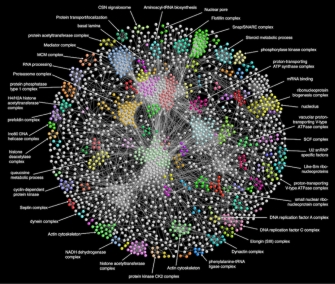
Determining the composition of protein complexes is an essential step toward understanding protein function and the cell as an integrated system. Using co-affinity purification coupled to mass spectrometry analysis, we examined protein associations for thousands of individual FLAG-HA epitope-tagged Drosophila proteins. Analysis of this data using a statistical framework designed to define individual protein-protein interactions led first to the generation of a partial Drosophila protein map encompassing 556 protein complexes (see Guruharsha, et al., 2011). This is a large project involving many collaborators and we are now in the final stages of assembling a new more complete map of the Drosophila proteome (DPiM : Drosophila Protein interaction map), based on the analysis of more than 10,000 individual epitope-tagged proteins. Importantly, given the now published human proteome network led by the Gygi and Harper laboratories, we are in a unique position to make extensive comparisons between the protein interactions in the fly vs. the human. Moreover, the DPiM defines potential novel members for several important protein complexes and assigns functional links to hundreds of protein-coding genes lacking previous experimental annotation.
Guruharsha, K.G., Rual, J.F., Zhai, B., Mintseris, J., Vaidya, P., Vaidya, N., Beekman, C., Wong, C., Rhee, D.Y., Cenaj, O., McKillip, E., Shah, S., Stapleton, M., Wan, K.H., Yu, C., Parsa, B., Carlson, J.W., Chen, X., Kapadia, B., VijayRaghavan, K., Gygi, S.P., Celniker, S.E., Obar, R.A., and Artavanis-Tsakonas, S. (2011). A protein complex network of Drosophila melanogaster. Cell. 147(3):690-703.